Keratoconus is a progressive ectatic disorder of the cornea characterized by thinning of the central cornea that leads to protrusion and progressive, irregular astigmatism. The etiopathogenesis of keratoconus is still being investigated; it may be the final manifestation of diverse pathologic processes. Better understanding of the disease, newer imaging modalities, and advances in refractive surgery have allowed earlier and accurate diagnosis of keratoconus.
The prevalence of keratoconus varies widely (0.0003% to 2.3%) depending on the geographic location, diagnostic criteria, and patient population.1,2 Environmental factors such as sunshine and hot weather are associated with a higher prevalence of keratoconus, and UV–light-induced oxidative stress may play a role a play as well.3 Ethnic differences may account for a greater prevalence of keratoconus in Asian (Indian, Pakistani, and Bangladeshi) individuals compared with whites.4,5 Family history of keratoconus has been found to be variable. The US Collaborative Longitudinal Evaluation of Keratoconus study reported a rate of 13.5%, whereas a study in Israel, where the prevalence is high, reported a rate of 21.74%.6
Keratoconus affects people of both sexes, although it is unclear whether significant differences between males and females exist. Eye-rubbing associated with atopy, ocular allergies, Down syndrome, and tapetoretinal degenerations may be linked to a higher incidence of keratoconus.7 Although most often seen bilaterally, asymmetry is possible. Unilateral cases of keratoconus do exist; however, their frequency, if appropriate diagnostic criteria and accurate examination techniques are used, are likely even lower than reported.8,9
To comprehend the etiopathogenesis of keratoconus, it is important that the various mechanisms that contribute to it are understood.
GENETIC PREDISPOSITION
A genetic basis for keratoconus has been an accepted theory considering its familial occurrence and high concordance in monozygotic twins. Kaya et al10 found the incidence of keratoconus to be 11% in first-degree relatives of patients with the disease, as opposed to a 0.05% incidence of keratoconus in the general population.
Studies have identified multiple genetic loci. In a Japanese study, HLA-A26, B40, and DR9 were associated with keratoconus in younger individuals.11 Udar et al12 demonstrated the uniqueness of a 7-base deletion in intron 2 of the SOD1 gene to the keratoconus phenotype. Héon et al13 identified mutations in the visual system homeobox (VSX) 1 gene for two distinct inherited corneal dystrophies and one mutation (R166W) responsible for keratoconus that altered the homeodomain and impaired the DNA binding. However, in another study,14 sequencing the coding region of VSX2 did not reveal a sequence variant segregating with disease in any of the families studied; this was the first report linking keratoconus to 14q24.3.
We have studied (unpublished data) the clinical features and VSX1 gene in 20 patients of eight families from India. In the coding region of VSX1, one novel missense heterozygous mutation (L268H) was identified in two unrelated keratoconus patients. Another family of three members had a novel heterozygous missense substitution (S251T). These variants cosegregated with the disease phenotype in all affected individuals but were not detected in the unaffected family members and 50 other healthy control patients. In silico analysis revealed that L268H is a pathogenic variant affecting the protein coded by VSX1 and that S251T had a neutral effect on the functional properties of VSX1.
In another study (unpublished data), we examined a trio of patients from a consanguineous Indian family with pellucid corneal marginal degeneration or keratoconus using homozygosity mapping. The regions of significant homozygosity in this family were on chromosomes 12p13.1 and 14q11.2, and significant SNPs in the region were rs1544671 and rs3811259, respectively. In a keratoconus linkage replication study, Bisceglia et al15 showed that chromosomal regions 5q32-q33, 5q21.2, 14q11.2, and 15q2.32 exhibited the strongest evidence of linkage. Equal evidence of linkage was also detected to both 1p36.23-36.21 and 8q13.1-q21.11, suggesting a digenic linkage.16
Also promising is a gene encoding the enzyme lysyl oxidase (LOX), located under a linkage peak at 5q23.2. LOX is a copper-dependent amine oxidase that is responsible for generating lysine-derived crosslinks in collagen and elastin.17 A change in LOX expression or the activity of its product may cause a decrease in crosslinking of collagen fibers, resulting in biomechanical weakening of the cornea and contributing to keratoconus development.
AT A GLANCE
• A better understanding of keratoconus, newer
imaging modalities, and advances in refractive surgery
have allowed earlier and more accurate diagnosis.
• Genetic, immunologic, and biochemical changes may
play important roles in keratoconus pathophysiology
and should be considered in prevention, diagnosis,
prognosis, and therapy.
• Keratoconus is likely to be a multifactorial,
multiagency disorder with varied inheritance patterns
and with environmental factors playing a role in the
causation of the disease.
We analyzed the expression of LOX (common variants LOXL1/LOXL2) and collagen (collagen variant 1A1 and variant 4A1) and found that the LOX transcript levels are reduced in keratoconus patients compared with controls.18 However, it is yet to be determined how LOX activity is altered in keratoconus and whether its reduction contributes to the pathogenesis of this progressive disorder. Further, it is not yet known if deregulation of LOX expression levels in the corneal epithelium and its activity correlate with disease severity and expression of other inflammatory genes known to be involved in keratoconus.
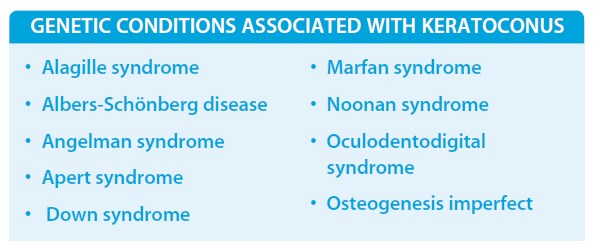
Keratoconus has also been associated with several genetic conditions, further reiterating that the disease can have a genetic predisposition.
INFLAMMATORY AND IMMUNOLOGIC ETIOLOGY
Keratoconus is traditionally considered a noninflammatory disease. However, several studies of tear film, serum cytokines, and tear proteomics conducted in the past decade have indicated an association of keratoconus with the expression of inflammatory mediators, such as proinflammatory cytokines, cell adhesion molecules and matrix metalloproteinases.19-26
The interleukin 1 (IL1) family is a group of 11 cytokines that plays a central role in the regulation of immune and inflammatory responses to infections or sterile insults. It has been found that polymorphisms in the IL1-α and IL1-β gene regions are associated with keratoconus in Korean and Japanese populations.19 Because IL1-α is upregulated in corneal trauma and inflammation, keratoconic corneas may muster an insufficient oxidative defense under such conditions.20 One study found higher levels of IL6, tumor necrosis factor alpha (TNF-α), and matrix metalloproteinase 9 (MMP9) in keratoconus eyes, and MMP9 was found to be the only independent variable associated with the degree of keratoconus.21,22
The MMPs are a family of zinc and calcium ion-dependent enzymes of differing substrate specificity but similar structure and catalytic mechanism. Physiologically, these enzymes participate in tissue remodeling. Pathologically, they cause disruption or disintegration of extracellular matrices. It has been shown that the tears of patients with anterior segment diseases induced by systemic disease or those that induce an inflammatory response contain MMP9. The main source of these enzymes appears to be the granulocyte fraction of white blood cells, which accumulate at sites of ocular inflammation.23
We have evaluated the levels of MMP9 in tears of Indian patients and investigated whether its regulation depends on proinflammatory cytokines, IL6, and TNF-α. We also evaluated the effects of the immunomodulator cyclosporine A (CyA) on cultured corneal epithelial cells and in patients with keratoconus. We found that MMP9, TNF-α, and IL6 levels were upregulated at the mRNA level in the epithelia of keratoconus patients. Similarly, tears collected from keratoconus patients exhibited high levels of MMP9 and IL6 protein, which further correlated with disease progression. However, the observed correlation with TNF-α in in vitro culture was not reflected in patient tears. CyA treatment significantly reduced the mRNA expression levels of IL6 and TNF-α in short- and long-term treatments but reduced MMP9 levels only in long-term treatment in corneal epithelial cell cultures. Further, treatment of keratoconus patients with CyA for about 6 months reduced tear MMP9 levels and led to a local reduction in corneal curvature as determined by corneal topography.24
Keratoconus is associated with several autoimmune and allergic conditions.25 This further supports the importance of the immune system in the pathogenesis of keratoconus. There is also a strong association of keratoconus with contact lens use, presumably due to the release of inflammatory mediators secondary to the mechanical stimulation of the corneal surface. Studies have shown that wearing rigid gas-permeable contact lenses induces overexpression of IL6, TNF-α, intercellular adhesion molecule 1 (ICAM-1), and vascular cell adhesion molecule 1 (VCAM-1) in the tears of patients with keratoconus.26
PROTEOMICS IN KERATOCONUS
Tissue extracts or fluids have revealed interesting protein profiles and signaling networks that are useful as clinical biomarkers for disease progression and as potential modes of therapeutic intervention. Analysis of differential spots has identified gelsolin, S100A4, and cytokeratin 3 to be highly overexpressed in keratoconus epithelium and alpha enolase to be slightly upregulated.27 A subsequent study using the same strategy found alpha enolase and beta actin to be poorly expressed in corneal wing and superficial epithelial cells from keratoconus patients.
Studies have shown more than 1,500 proteins and peptides in the tear film, in addition to lipids, cytokines, small molecules, and metabolites.28 These tear components consist largely of lysozyme, serum albumin, lactoferrin, secretory immunoglobulin A, proline-rich proteins, tear lipocalin, and lipophilin.29 While one study found immunoglobulin alpha and kappa, lipocalin, lysozyme C, and precursors to prolactin to be associated with keratoconus,30 another study of tear film proteomics found reduced expression of a few novel proteins, zinc-α2-glycoprotein, and immunoglobulin kappa chain, as well as lactoferrin.31 A recent study showed that keratoconus tears contained significantly reduced levels (by approximately 50%) of total protein. It also found that levels of lactoferrin and secretory immunoglobulin A were reduced in keratoconus tears and negatively correlated with keratometry readings.32 However, none of the studies have been able to produce stable biomarkers useful for clinical stratification of patients.
We studied (unpublished data) the quantitative tear proteomic profile of an Indian cohort of keratoconus patients using tandem liquid chromatography-mass spectrometry/mass spectrometry techniques. In total, the tear proteome from keratoconic eyes and controls was about 300 to 400 proteins in more than 50% of cases, with a cutoff for false discovery rate at 1%. About 48 proteins were identified to be present in every sample and more than 1,000 proteins detected in at least three samples in the cohort. Further, there was significant deregulation of several proteins in keratoconic eyes, with many showing an increase in average levels across the patient cohort corresponding to clinical grades, particularly lipocalin 1; phospholipase A2, group II, isoform A; secretoglobin family 2A member 1; mesothelin; and crystallin alphaB. These proteins therefore demonstrate the potential to be considered as biomarkers for predicting the progression of keratoconus.
A recent study showed that keratoconus tears contained significantly reduced levels (by approximately 50%) of total protein.
CONCLUSION
Genetic, immunologic, and biochemical changes may play important roles in keratoconus pathophysiology. These changes may be considered in the prevention, diagnosis, prognosis, and therapy of the disease. Keratoconus is likely to be a multifactorial, multiagency disorder with varied inheritance patterns and with environmental factors playing a role in the causation of the disease. n
1. Gorskova EN, Sevost'ianov EN. Epidemiology of keratoconus in the Urals. Vestn Oftalmol. 1998;114:38-40.
2. Jonas JB, Nangia V, Matin A, Kulkarni M, Bhojwani K. Prevalence and associations of keratoconus in rural Maharashtra in central India: The Central India Eye Medical Study. Am J Ophthalmol. 2009;148:760-765.
3. Gokhale NS. Epidemiology of keratoconus. Indian J Ophthalmol. 2013;61:382-383.
4. Georgiou T, Funnell CL, Cassels-Brown A, O'Connor R. Influence of ethnic origin on the incidence of keratoconus and associated atopic diseases in Asian and white patients. Eye (Lond). 2004;18:379-383.
5. Pearson AR, Soneji B, Sarvananthan N, Sandforth-Smith JH. Does ethnic origin influence the incidence or severity of keratoconus? Eye (Lond). 2000;14:625-628.
6. Zadnik K, Barr JT, Edrington TB, et al. Baseline findings in the Collaborative Longitudinal Evaluation of Keratoconus (CLEK) study. Invest Ophthalmol Vis Sci. 1998;39:2537-2546.
7. Krachmer JH, Feder RS, Belin MW. Keratoconus and related non-inflammatory corneal thinning disorders. Surv Ophthalmol. 1984;28:293-322.
8. Amsler M. Some data on the problem of keratoconus. Bull Soc Belge Ophthalmol. 1961;129:331-354.
9. Chopra I, Jain AK. Between eye asymmetry in keratoconus in an Indian population. Clin Exp Optom. 2005;88:146-152.
10. Kaya V, Utine CA, Altunsoy M, Oral D, Yilmaz OF. Evaluation of corneal topography with Orbscan II in first-degree relatives of patients with keratoconus. Cornea. 2008;27(5):531-534.
11. Adachi W, Mitsuishi Y, Terai K, et al. The association of HLA with young-onset keratoconus in Japan. Am J Ophthalmol. 2002;133(4):557-559.
12. Udar N, Atilano SR, Small K, Nesburn AB, Kenney MC. SOD1 haplotypes in familial keratoconus. Cornea. 2009;28(8):902-907.
13. Héon E, Greenberg A, Kopp KK, et al. VSX1: a gene for posterior polymorphous dystrophy and keratoconus. Hum Mol Genet. 2002;11(9):1029-1036.
14. Liskova P, Hysi PG, Waseem N, Ebenezer ND, Bhattacharya SS, Tuft SJ. Evidence for keratoconus susceptibility locus on chromosome 14: a genome-wide linkage screen using single-nucleotide polymorphism markers. Arch Ophthalmol. 2010;128(9):1191-1195.
15. Bisceglia L, De Bonis P, Pizzicoli C, et al. Linkage analysis in keratoconus: replication of locus 5q21.2 and identification of other suggestive Loci. Invest Ophthalmol Vis Sci. 2009;50(3):1081-1086.
16. Burdon KP, Coster DJ, Charlesworth JC, et al. Apparent autosomal dominant keratoconus in a large Australian pedigree accounted for by digenic inheritance of two novel loci. Hum Genet. 2008;124(4):379-386.
17. Bykhovskaya Y, Li X, Epifantseva I, et al. Variation in the lysyl oxidase (LOX) gene is associated with keratoconus in family-based and case-control studies. Invest Ophthalmol Vis Sci. 2012;53(7):4152-4157.
18. Mikami T, Meguro A, Teshigawara T, et al. Interleukin 1 beta promoter polymorphism is associated with keratoconus in a Japanese population. Mol Vis. 2013;19:845-851.
19. Shetty R, Reshma AR, Arunapriya S, et al. Attenuation of lysyl oxidase and collagen gene expression in keratoconus patient corneal epithelium corresponds to disease severity. Mol Vis. 2014. Accepted for publication.
20. Olofsson EM, Marklund SL, Pedrosa-Domellöf F, Behndig A. Interleukin-1alpha downregulates extracellular-superoxide dismutase in human corneal keratoconus stromal cells. Mol Vis. 2007;13:1285-1290.
21. Lema I, Durán JA. Inflammatory molecules in the tears of patients with keratoconus. Ophthalmology. 2005;112(4):654-659.
22. Lema I, Sobrino T, Durán JA, Brea D, Díez-Feijoo E. Subclinical keratoconus and inflammatory molecules from tears. Br J Ophthalmol. 2009;93(6):820-824.
23. Smith VA, Rishmawi H, Hussein H, Easty DL. Tear film MMP accumulation and corneal disease. Br J Ophthalmol. 2001;85(2):147-53.
24. Shetty R, Ghosh A, Lim RR, et al. Elevated expression of matrix metalloproteinase 9 and inflammatory cytokines in keratoconus patients is inhibited by cyclosporine A. Invest Ophthalmol Vis Sci. 2014. Accepted for publication.
25. Nemet AY, Vinker S, Bahar I, Kaiserman I. The association of keratoconus with immune disorders. Cornea. 2010;29(11):1261-1264.
26. Lema I, Durán JA, Ruiz C, Díez-Feijoo E, Acera A, Merayo J. Inflammatory response to contact lenses in patients with keratoconus compared with myopic subjects. Cornea. 2008;27(7):758-763.
27. Nielsen K, Vorum H, Fagerholm P, et al. Proteome profiling of corneal epithelium and identification of marker proteins for keratoconus, a pilot study. Exp Eye Res. 2006;82:201-209.
28. Zhou L, Zhao SZ, Koh SK, et al. In-depth analysis of the human tear proteome. J Proteomics. 2012;75:3877-3885.
29. de Souza GA, Godoy LM, Mann M. Identification of 491 proteins in the tear fluid proteome reveals a large number of proteases and protease inhibitors. Genome Biol. 2006;7:R72.
30. Pannebaker C, Chandler HL, Nichols JJ. Tear proteomics in keratoconus. Mol Vis. 2010;16:1949-1957.
31. Lema I, Brea D, Rodriguez-Gonzalez R, Diez-Feijoo E, Sobrino T. Proteomic analysis of the tear film in patients with keratoconus. Mol Vis. 2010;16:2055-2061.
32. Balasubramanian SA, Pye DC, Willcox MD. Levels of lactoferrin, secretory IgA and serum albumin in the tear film of people with keratoconus. Exp Eye Res. 2012;96:132-137.
Chaitra Jayadev, DOMS
- Consultant at Narayana Nethralaya
- drchaitra@hotmail.com
- Financial disclosure: None
Rohit Shetty, FRCS, DNB
- Senior Consultant and Vice Chairman of Narayana Nethralaya
- drrohitshetty@yahoo.com
- Financial disclosure: None






